As Qingming Festival approaches, barley in fields enters the critical heading and booting stage. As an important raw material for grain, forage and beer brewing in China, high yield and stress resistance of barley are expected in spring farming, and also key topics in agricultural genomics research.
iGeneTech has long been engaged in plant and animal whole exome sequencing. Based on its independently developed IGT® Oligo Pools synthesis platform and TargetSeq® technology platform, the company has built a product matrix covering nearly 30 core species. Among them, the Barley Whole Exome Solution features high uniformity and high specificity, providing reliable tools for genomic research and molecular breeding.

Technical Choice for Large Genomes
The genome of barley (Hordeum vulgare) is approximately 5 Gb. The current high-quality reference genome Morex V3 has assembled about 4.2 Gb, consisting of approximately 80% repetitive sequences. It has a complex structure and relatively sparse distribution of functional genes.
Against this background, direct whole-genome sequencing (WGS) often faces:
· High sequencing cost
· High analysis difficulty
· Limited efficiency in functional variant interpretation
In contrast, whole exome capture technology focuses on coding regions (CDS), enabling:
· Precise enrichment of functional variant regions
· Significantly reduced sequencing and analysis costs
· Improved efficiency of key gene interpretation
Related Research
Case 1 | Exome sequencing facilitates comparative genomics and population genetics of wheat and barley (2025)
Researchers performed whole exome sequencing and population genetic analysis on 672 wheat accessions and 679 barley accessions. They constructed the ancestral Triticeae karyotype, systematically revealed the convergent selection patterns of wheat and barley in domestication, stress resistance, yield and other traits, identified numerous conserved homologous genes and key variant loci, and proposed a new strategy for cross-crop translational breeding. This provides important genetic resources and theoretical support for precision breeding of cereal crops.
Case 2 | Barley exome capture assists fine mapping and identification of key candidate genes (2025)
In a study of quantitative trait locus (QTL) and candidate gene mapping in two-row spring barley, the research team used exome capture sequencing data to increase marker density for finer QTL mapping and candidate gene discovery. In 31.65% of the QTLs, markers in exonic regions showed stronger association signals than the original 50k SNP chip markers, enabling more accurate identification of candidate genes.
iGeneTech Barley Whole Exome Solution
iGeneTech Barley Whole Exome Panel is designed based on the 37.1 Mb CDS region of the reference genome Morex V3 (GCA_904849725.1), with 466,714 unique probes, enabling efficient and uniform capture of nearly 30,000 barley coding genes.
Paired with the TargetSeq One® V3.0 hybrid capture system, the capture efficiency of barley whole exome exceeds 70%, and 50×+ effective depth can be achieved with 5 Gb of sequencing data.
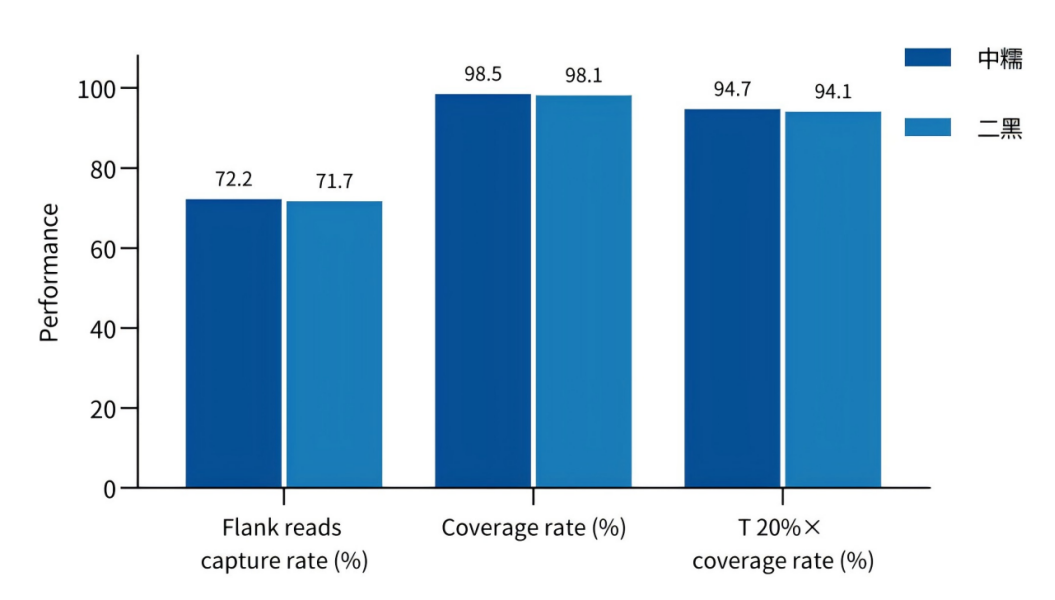
Figure 1 Data performance of the Barley Whole Exome Panel across different barley varieties.
Product Advantages
· High Specificity Focuses on the whole exome region and directly obtains sequence information of gene CDS regions; measured coverage rate exceeds 98%.
· Cost-Effective Compared with whole-genome sequencing (WGS), whole exome sequencing has lower cost while achieving high depth and accuracy.
· Flexible Customization Based on the self-developed massive parallel synthesis platform, genome-wide background loci or functional loci can be added as required.
iGeneTech Barley Whole Exome Solution is widely used in diverse scenarios, supporting large-scale population studies, mutant libraries, gene mapping, genetic breeding and other research needs, providing precise technical support for scientific breakthroughs in related fields.
Related Product List
Product Name | Specification (rxn) | Cat. No. |
Barley Whole Exome Panel | 24 / 96 | PH2002775 / PH2002772 |
TargetSeq One® Hyb & Wash Kit v3.0 (for Illumina) | 24 / 96 | C11534 / C11532 |
IGT® Enzyme Plus Library Prep Kit V3 | 96 rxn | C11112 |
IGT® Adapter & 10 nt UDI Primer 1–768 (for Illumina, plate) | 768×1 rxn | C11282 |
References
[1] Author Correction: Striking convergent selection history of wheat and barley and its potential for breeding. Nature Plants, 2025, 11(12): 2581. DOI: 10.1038/s41477-025-02182-8.
[2] New loci and candidate genes in spring two-rowed barley detected through meta-analysis of a fieldtrial European network. Theoretical and Applied Genetics, 2025, 138(7). DOI: 10.1007/s00122-025-04934-8.

 CN
CN


