Application Background
In the field of genetic disease testing, Whole Exome Sequencing (WES) has become an important first-line method. Compared with single-gene testing or disease panels, WES covers more than 20,000 gene coding regions in a single test, providing comprehensive genetic information for rare disease diagnosis.
However, in practical testing, researchers and clinicians often face difficulties in sequencing high-GC regions. These regions suffer from low amplification efficiency, uneven library distribution, and unstable signals in sequencing regions, leading to missed detections and inaccurate data in key areas.
Table 1 List of partially high-GC and complex hard-to-detect sequences of clinical concern
Region | Clinical Significance | Common Sequencing Issues |
TERT Promoter | Key region for oncological and genetic testing | High GC content, concentrated hotspot loci; insufficient sequencing depth and poor locus interpretability |
MECP2 Exon1 | Pathogenic variants related to Rett syndrome may locate in exon 1 | High GC content; risk of insufficient coverage or missed detection |
CEBPA Coding region | Definite pathogenic gene for familial AML | Single exon with high GC content; difficult amplification, easily affecting the integrity of variant detection |
SHANK3 Coding region | Clearly associated with neurodevelopmental abnormalities, ASD and Phelan-McDermid syndrome | Elevated overall GC content in coding region; complex detection, unstable capture and sequencing performance |
RPGR ORF15 | Critical region for retina-related diseases | Complex and hard-to-sequence region; insufficient coverage and difficult alignment |
High-GC regions in whole exome sequencing are usually limited by three factors:
· Insufficient probe coverage
· Restricted hybrid capture
· Sequencing bias
iGeneTech relies on its proprietary liquid-phase chip capture technology, using denser probes for high-GC regions, optimized probe layout, and improved hybrid capture reagent systems to enhance capture efficiency in complex regions. GeneMind has broken through difficult sequencing regions with its CMS sequencing technology, further improving performance in high-GC regions and reducing coverage gaps.
As a result, iGeneTech’s whole exome products are more competitive in detecting high-GC and complex regions.

AIExome Series: Full-Process Optimization for High-GC Regions
High-GC Region Probe Design — The Source of Accuracy
High-GC regions form more stable secondary structures, and conventional probe design often suffers from reduced binding efficiency and uneven capture. To address this, iGeneTech uses a targeted probe layout strategy: denser coverage for GC-abnormal regions, optimized probe distribution, and improved uniformity within complex regions, maximizing capture performance at the source.
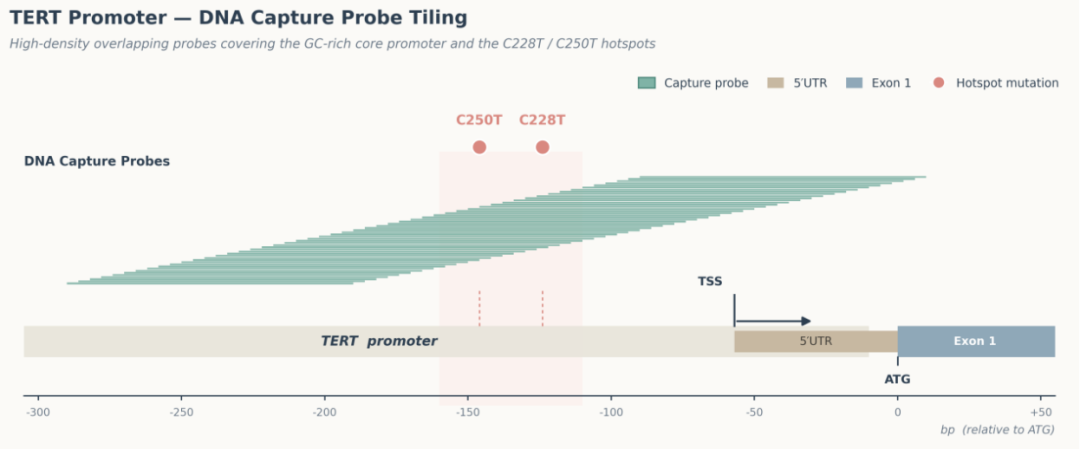
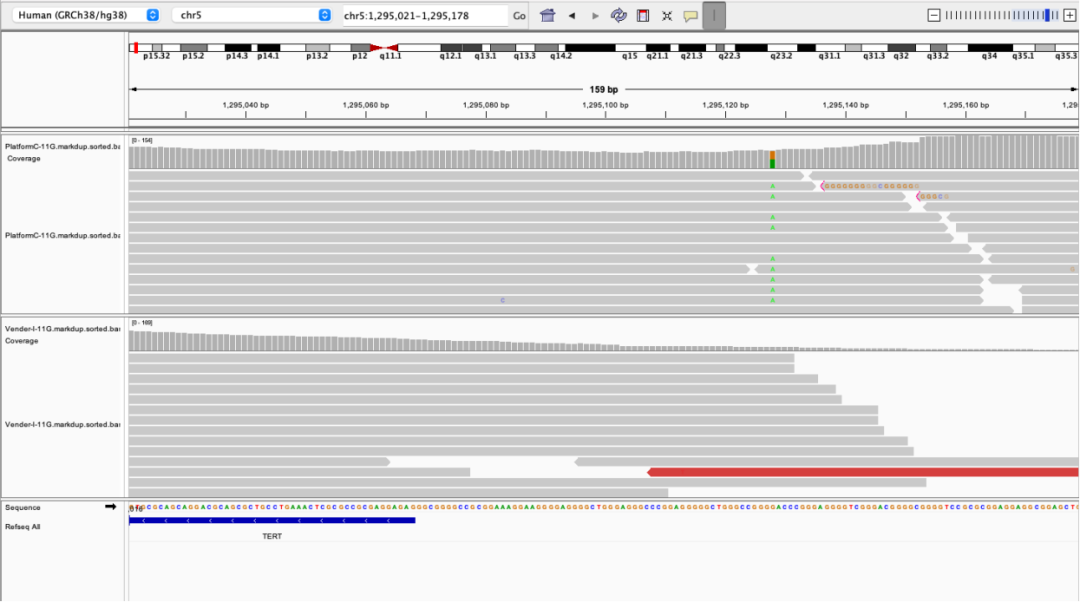
Figure 1 Probe Design and Actual Sequencing Data Comparison of TERT Promoter Region for AIExome V5 Core Edition
The displayed data compare the coverage depth of the TERT promoter region across different whole-exome sequencing products under the same sequencing platform and identical data volume. For two key mutation loci in this region, the sequencing depth of AIExome V5 Core Edition reaches 97× and 90× respectively, while the competitor’s sequencing depth is only 25× and 18× correspondingly.
Hybrid Capture System — The Core of Performance
In the hybridization system, high-GC templates tend to form stable paired structures, hindering effective binding between probes and target fragments. Thus, optimization of hybridization conditions is critical. iGeneTech adjusts hybridization temperature, salt ion conditions, reaction components, and washing conditions, greatly improving data uniformity and significantly enhancing recovery efficiency and coverage in high-GC regions.
Across multiple sequencing platforms, GC-abnormal regions show excellent performance.
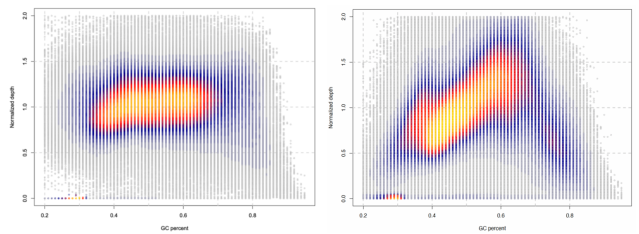
Figure 2 Balanced GC Coverage of AIExome V5 Core Edition
The horizontal axis represents regions with different GC contents, and the vertical axis indicates the normalized depth of each region. The left panel shows data from AIExome V5 Core Edition combined with Targetseq One® Hyb and Wash Kit V3.0 capture reagent, sequenced on the Illumina platform with PE150 mode; the right panel presents capture data of Axx V8, also sequenced on the Illumina platform with PE150 mode.
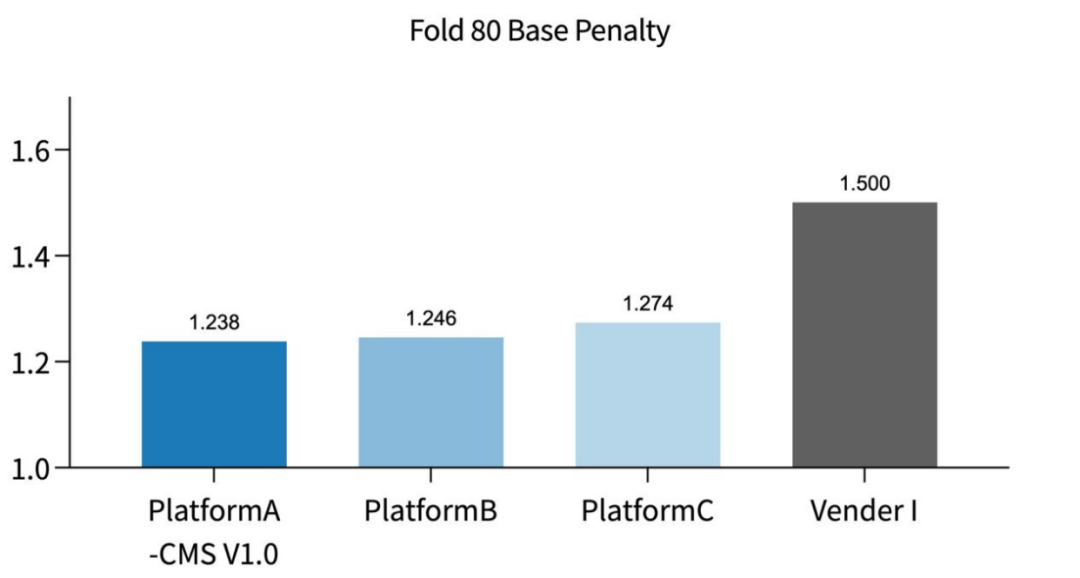
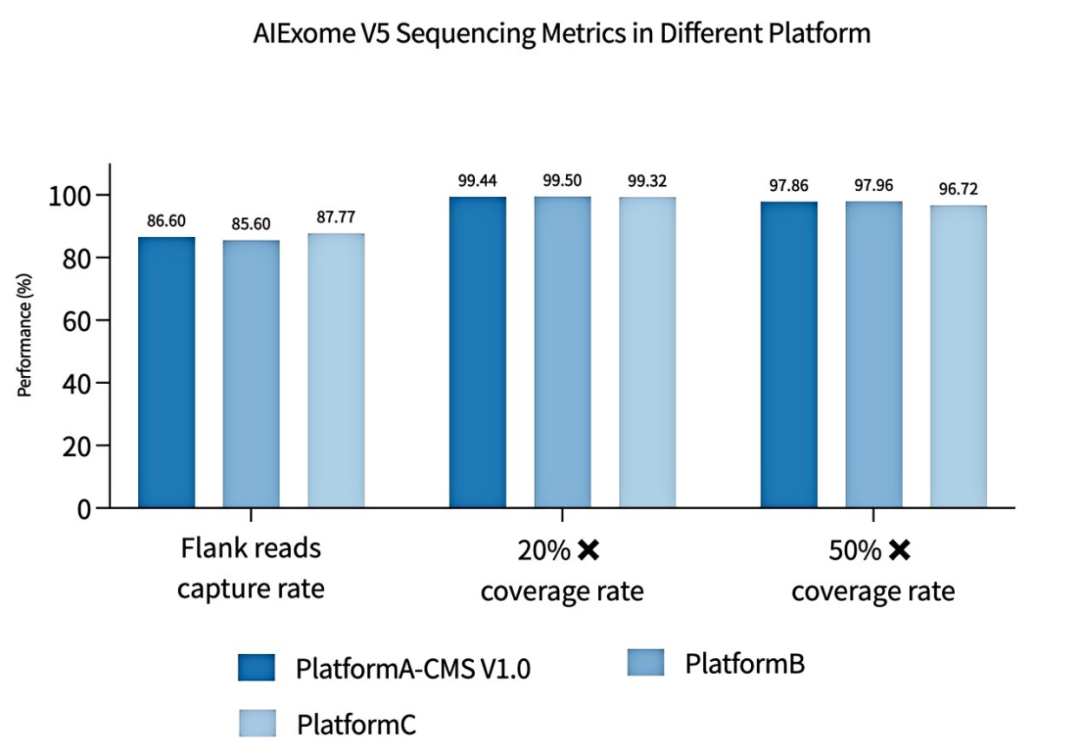
Figure 3 Excellent Sequencing Metrics and Uniformity of AIExome V5 Core Edition Across Different Platforms (Fold 80 Base Penalty)
The first three products show the Fold 80 Base Penalty metrics of AIExome V5 Core Edition on different sequencing platforms. The Fold 80 data of the competitor’s whole-exome product are sourced from its official promotional brochure.
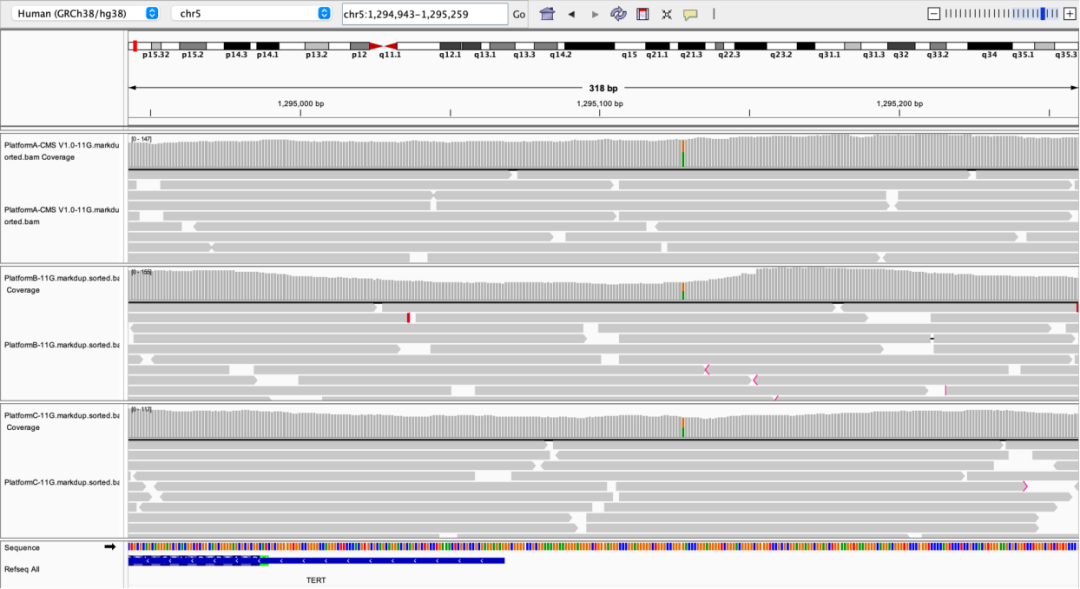
TERT promoter region
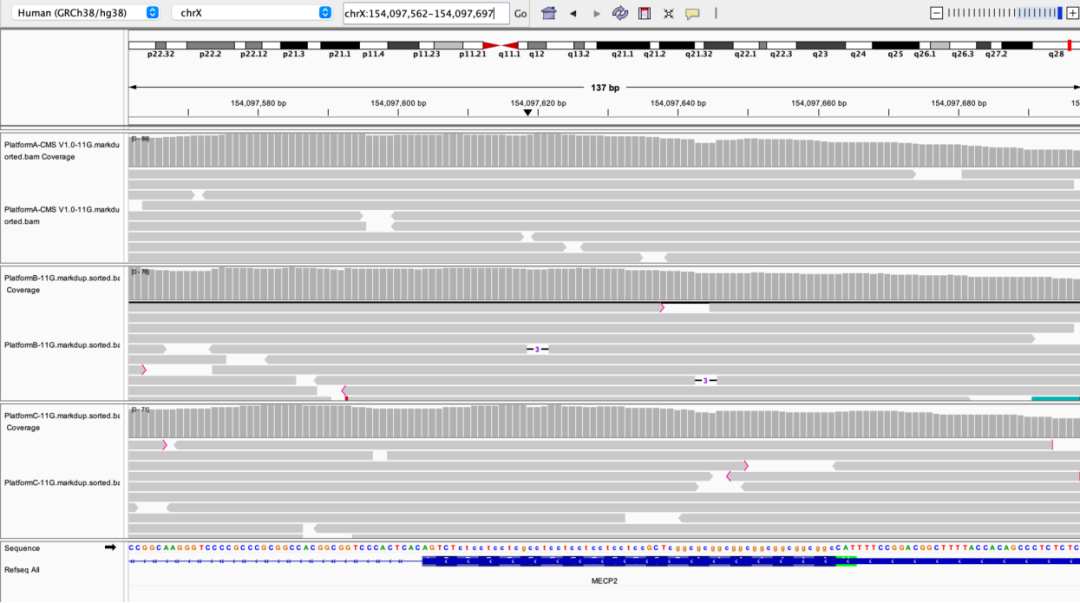
MECP2 Exon1
CEBPA
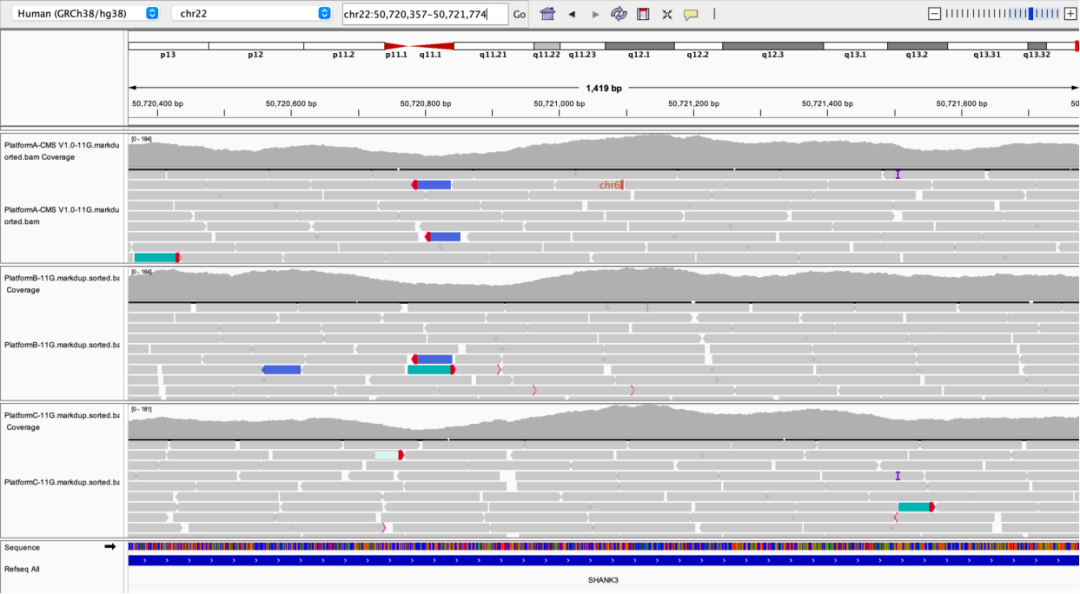
SHANK3 Exon24
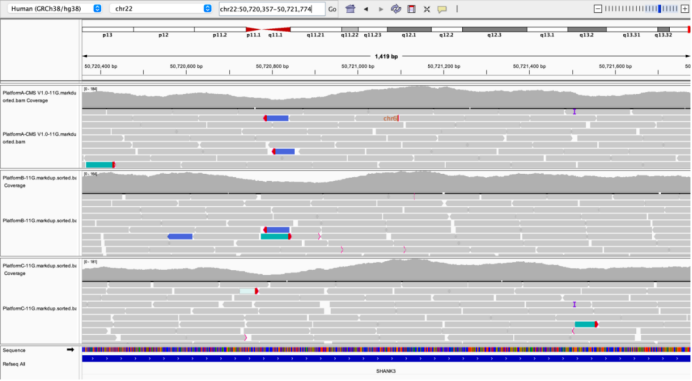
Figure 4 Outstanding performance of AIExome V5 Core Edition in key high-GC regions.
GeneMind: Crossing Mountains and Seas in Sequencing
In the sequencing stage, high-GC fragments still face challenges such as low amplification efficiency and difficult extension by sequencing polymerases.
Focusing on core bottlenecks in sequencing, GeneMind has launched CMS (Cross Mountains and Seas) sequencing technology, which has been industrialized as the CMS Sequencing Kit V1.0 on the SURFSeq 5000. It perfectly inherits the capture advantages of iGeneTech, significantly strengthens sequencing capability in genomic difficult regions, and delivers unbiased, high-precision data at the Q50 level (99.999% accuracy). It achieves breakthrough improvements especially in high-GC regions.
CMS-V1.0 shows the lowest error rate in raw sequencing accuracy.
All platforms adopted data with an average sequencing depth of 550× for analysis. After excluding polymorphic loci, each locus in the remaining target interval loci may have three types of single-base substitution errors. We counted the number of loci with a single-base error rate greater than or equal to a certain level. Among them, the eNPM (Error Numbers Per Million Positions) 01/03/05 of CMS V1.0 was significantly superior to the comparison platforms. Taking eNPM05 (the number of loci with an error rate ≥5% per million loci) as an example, the value of CMS was only 1/54 (20/1075) of PlatformB and 1/6 (20/118) of PlatformC.
Table 2 NPM (Numbers per Million Position) Statistics Across Different Platforms
Platform | p01 | p03 | p05 |
PlatformA-CMS V1.0 | 1,700 | 952 | 0 |
PlatformB | 24,781 | 3,014 | 1,075 |
PlatformC* | 6,754 | 451 | 118 |
* The low NPM performance of PlatformC results from the loss of a large number of low-coverage regions, which inherently have a high error rate.
GeneMind’s CMS sequencing reagent performs outstandingly in complex structural regions, significantly improving sequencing accuracy and coverage uniformity. It demonstrates superior data quality and stronger variant interpretation ability, especially in high-GC regions such as RPGR ORF15.
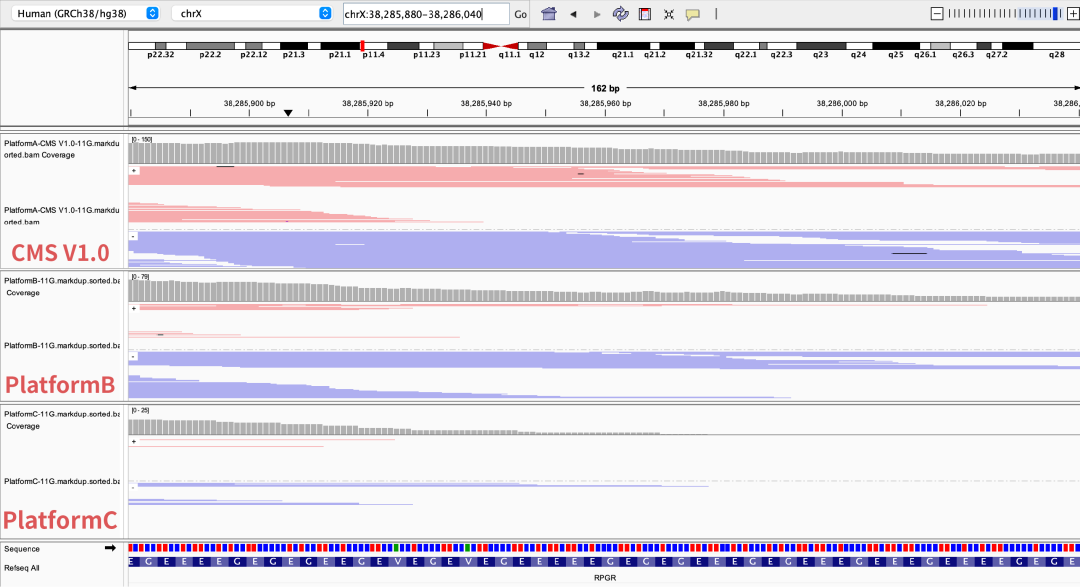
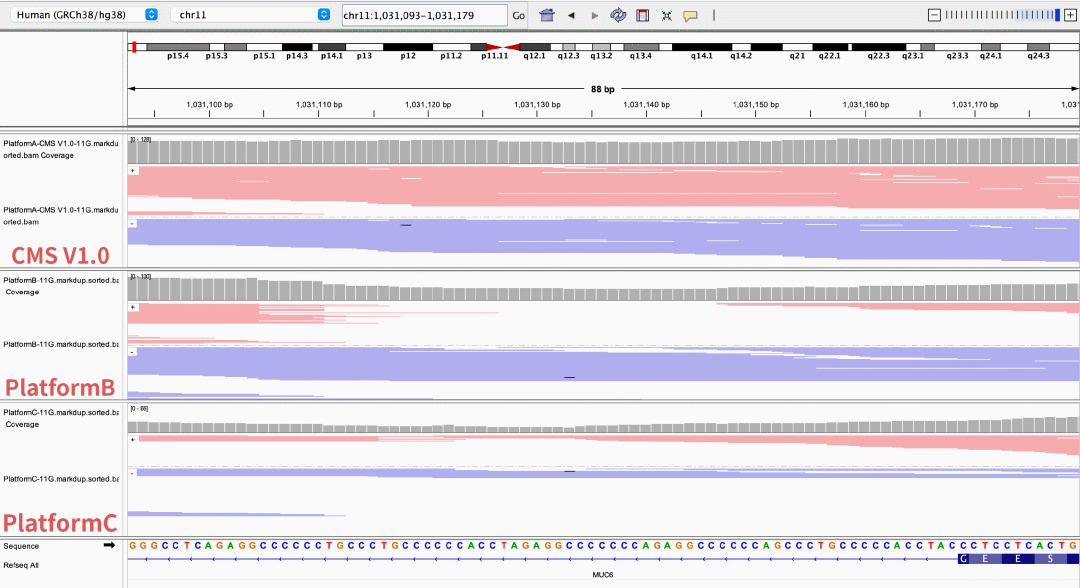
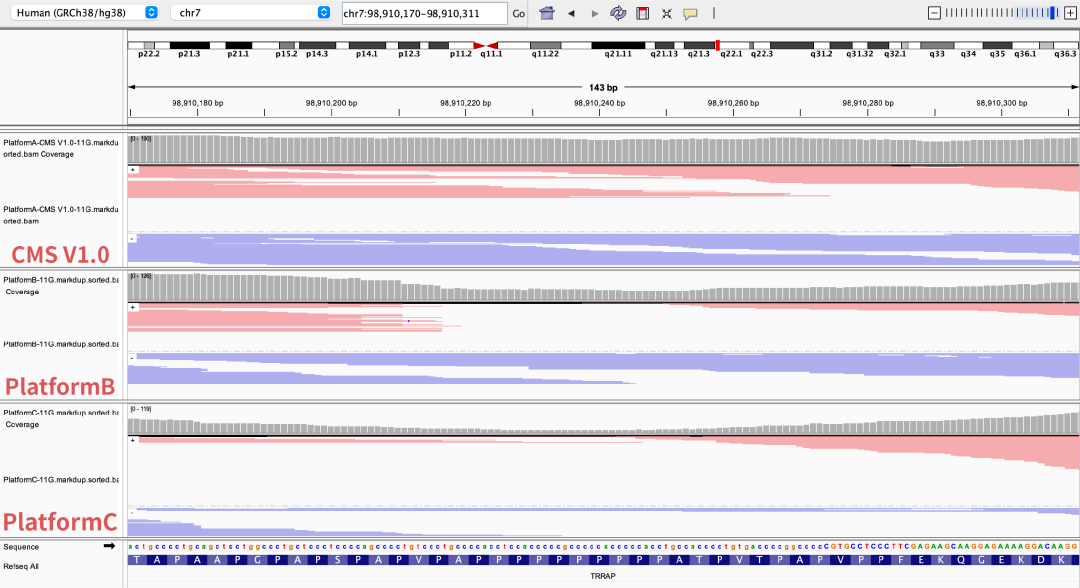
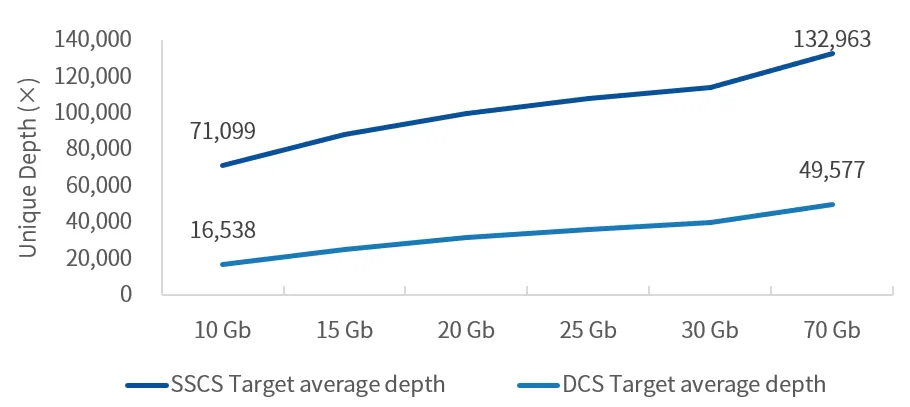
Figure 5 Coverage depth of partial high-GC regions on different sequencing platforms
The NA12878 sample was processed with enzymatic library construction, hybrid capture was performed using probes of AIExome V5 Core Edition, followed by sequencing on multiple platforms. All data were uniformly downsampled to a data volume of 11 Gb.
Practical Significance for Genetic Testing
For clinical genetic testing, regional coverage is fundamental, but the real value of testing lies in stable and consistent detection of key variant sites.
To evaluate consistency and stability in real-world testing, we performed capture with the AIExome V5 Core Panel using the NA12878 sample and further analyzed mutation sites.
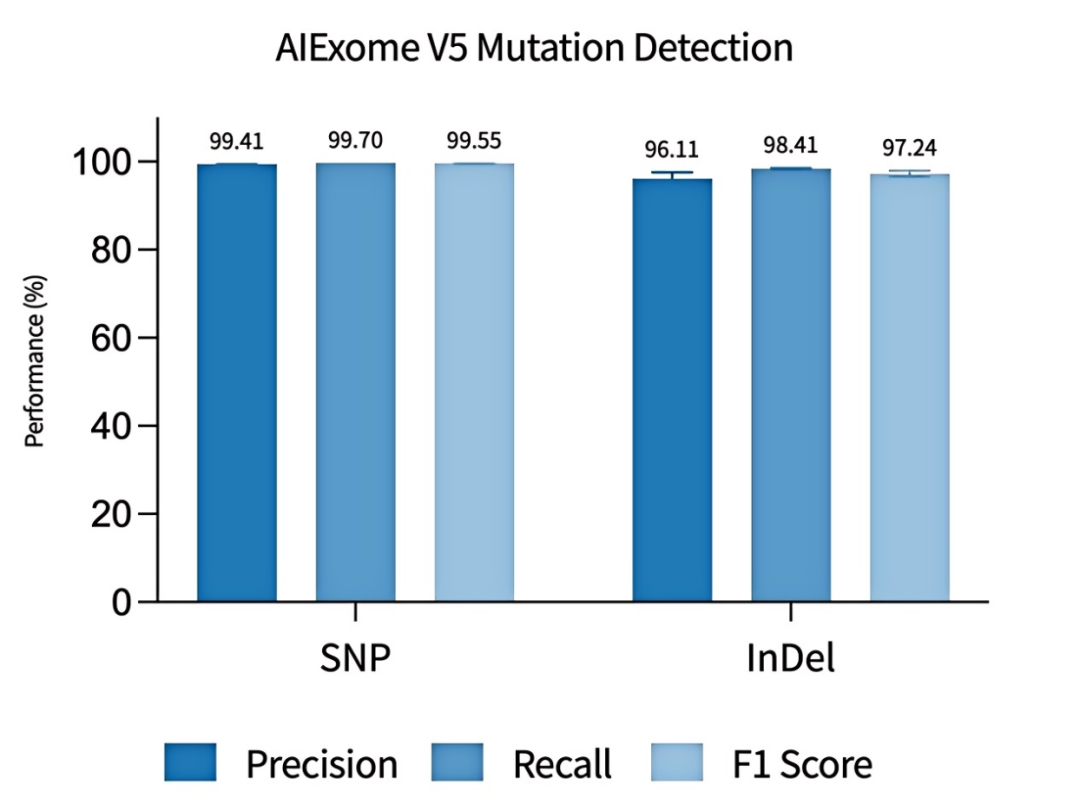
Figure 6 Evaluation of variant detection accuracy of AIExome V5 Core Edition
The NA12878 sample was adopted for enzymatic library construction, followed by hybrid capture with probes of AIExome V5 Core Edition and sequencing on multiple platforms. Data were uniformly downsampled to 11 Gb for variant concordance analysis. The corresponding variant VCF files and matched BED files of high-confidence regions were derived from the GIAB project, with the high-confidence version being NIST v3.3.2.
Conclusion
The collaboration between iGeneTech and GeneMind has achieved a true 1+1>2 effect of “capture + sequencing”. The CMS sequencing reagent is fully compatible with iGeneTech’s capture system without changing existing workflows, significantly enhancing sequencing capability in genomic difficult regions.
High-GC regions are no longer an insurmountable bottleneck for whole exome sequencing, supporting technological upgrades in rare disease diagnosis, early cancer screening, and other fields.
With domestic innovation, we make whole exome sequencing more comprehensive, stable, and reliable, and jointly promote the high-quality development of the genetic testing industry.

 CN
CN