Relapse remains the core challenge in the treatment of acute myeloid leukemia (AML). Measurable residual disease (MRD), the most robust independent prognostic marker for assessing remission depth and predicting relapse risk, demands standardized and precise detection in clinical practice.
AML MRD serves as a core index for prognosis evaluation and treatment guidance, with key clinical values:
· Precise prognostic stratification: Distinguish high/low relapse risk to guide treatment intensity adjustment.
· Treatment decision support: Optimize transplantation timing, preconditioning regimens and initiate targeted maintenance therapy.
· Early relapse warning: Detect residual lesions months before morphological relapse for timely intervention.
· Clinical trial endpoint: Accelerate new drug development and approval.
The 2025 AML MRD consensus released by the European Leukemia Network (ELN-DAVID) established a standardized diagnosis and treatment framework with 56 professional recommendations, advancing MRD detection toward ultra-high sensitivity, subtype stratification and precise clinical decision-making. MRD results are now stratified for treatment guidance instead of a single value; detection thresholds vary by technology—e.g., NPM1 detection threshold is reduced to 0.001%, ultra-high sensitivity NGS is recommended for FLT3-ITD, and molecular MRD monitoring should last 24 months post-transplantation.
AML MRD detection technologies evolve around three core goals: improving sensitivity, expanding coverage and guiding clinical practice precisely. From single-gene qPCR to multi-gene analysis and ultra-high sensitivity NGS, technological innovations continuously address clinical pain points and support precise AML diagnosis.
Comparison of AML MRD Detection Technologies
Technology |
Core Advantages
| Main Advantages
| Applicable Scenarios |
Multiparameter Flow Cytometry (MFC) | 90% patient applicability, easy operation, LSC detectability | Limited sensitivity (≈0.01%), high false positive rate, unable to track clonal evolution | AML patients without definite molecular targets; rapid preliminary MRD assessment |
qPCR/dPCR | High specificity, low cost, high standardization, absolute quantification | Narrow coverage (40%-60% patients), single target only | AML patients with definite molecular targets (e.g., NPM1, PML:RARA); low-cost rapid monitoring |
Ultra-high Sensitivity NGS | Ultra-high sensitivity (≤0.01%), multi-target simultaneous detection, customizable coverage for specific mutations (including rare ones) | High cost, strict DNA quality/input requirements (≥500ng), 3-7 day turnaround | High-risk AML patients (e.g., FLT3-ITD mutation); precise monitoring before/after transplantation; clinical research (e.g., new drug efficacy evaluation) |
iGeneTech AML MRD Solution
Designed based on hybridization capture sequencing, iGeneTech solution selects key AML-related mutation sites to detect multiple variants (base substitution, insertion/deletion, etc.). Combined with double-end Unique Molecular Identifier (UMI) technology, it enables flexible customization of coverage areas for efficient and sensitive detection.
1. Comprehensive Target Coverage
The AML MRD Panel targets 14 core genes with a 42.5 kb probe coverage area, accurately covering key markers such as NPM1 and FLT3. It detects multiple variant types, adapts to diverse clinical scenarios, and balances detection breadth and sequencing cost.
Core Genes Covered: ASXL1, CEBPA, CBL, EZH2, FLT3, IDH2, KIT, KRAS, NPM1, NRAS, RUNX1, U2AF1, TP53, IDH1.
2. Standardized Workflow
Equipped with full-process reagents (nucleic acid extraction, library construction, capture) and automated protocols, the solution features simplified reagents and workflows for stable and reliable results. It supports MGI, ILMN and Zonmagna sequencing platforms, with free MRD probe synthesis and 4-day delivery, plus a matched UMI bioinformatics analysis pipeline.
3. High-performance Capture
Using Shuimu Jiheng hematological tumor gDNA standard (MD030064) with 500 ng input, library construction was performed with enzyme digestion kit and double-end UMI adapters, followed by hybridization capture with all library products.
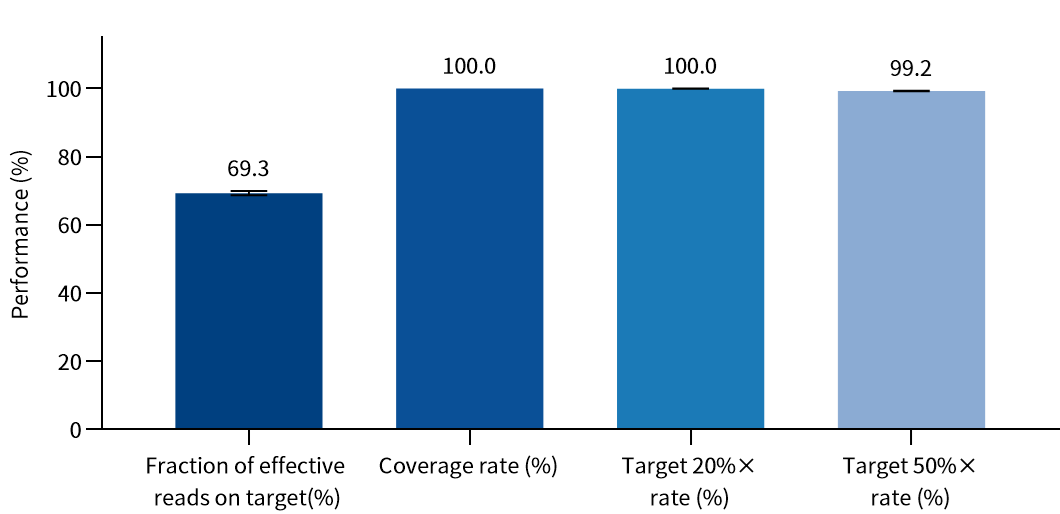
Figure 1. Capture Performance Test
4. Efficient Raw Molecule Recovery
MRD detection sensitivity depends on DNA input and final effective depth. More DNA input yields more raw molecules, but UMI-based deduplication and correction require sufficient sequencing depth to distinguish true mutations from sequencing errors, leading to increased sequencing data volume with higher DNA input.
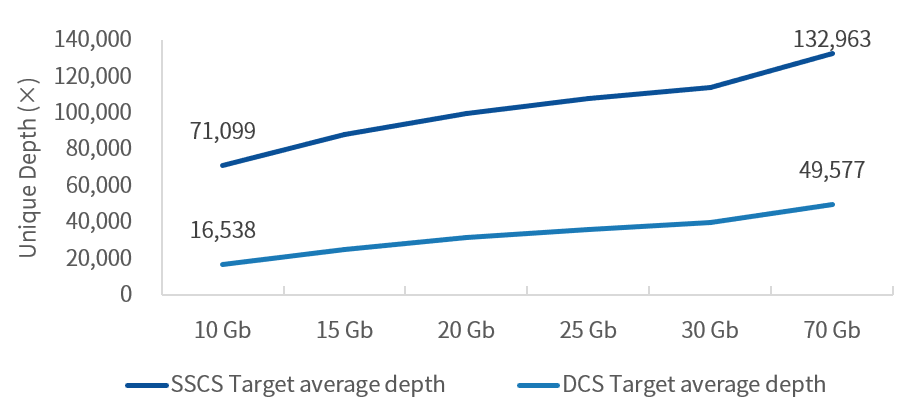
Figure 2. A test with 500 ng gDNA and 70 Gb high-depth sequencing (gradient random data truncation) via iGeneTech's UMI pipeline showed continuous growth of deduplicated depth in the 10~70 Gb range. SSCS (Single Strand Consensus Sequence) removes duplicates by UMI and genomic position; DCS (Duplex Consensus Sequence) corrects by matching sense/antisense strands .
Sensitivity and Data Volume by DNA Input (95% Detection Power)
Input (ng) | Raw Data (Gb) | DCS Mean Depth (x) | Detectable Frequency (1 Mutation) | Detectable Frequency (3 Mutations) |
200 | 4 | 6600 | 0.045% | 0.015% |
500* | 10* | 16500* | 0.018% | 0.006% |
1000 | 20 | 33000 | 0.009% | 0.003% |
1500 | 30 | 49500 | 0.006% | 0.002% |
*Measured data with 10 Gb raw bases at 500 ng input |
5. High-sensitivity Mutation Detection
· Gradient dilution of standard samples and detection with 10 Gb data volume showed the solution can detect ultra-low frequency mutations as low as 0.03%.
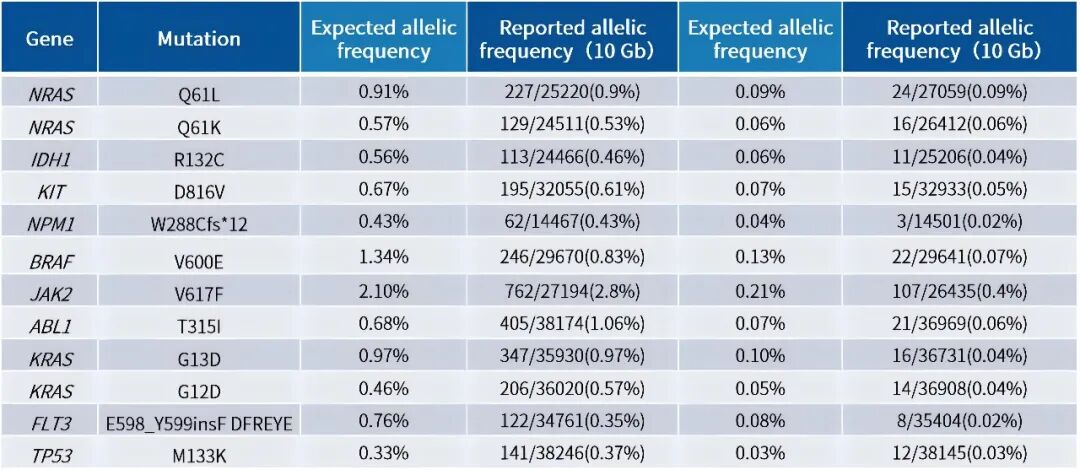
With the growing demand for standardized and personalized AML MRD detection, iGeneTech will continuously optimize its AML MRD solution. Based on professional R&D of upstream kits, the solution meets the large-scale, standardized testing needs of third-party clinical laboratories and the precise, customized needs of hospital-based research, supporting the clinical application and scientific exploration of AML MRD detection with reliable technical tools.
Reference
1. Cloos J, Valk PJM, Thiede C, et al. 2025 Update on MRD in Acute Myeloid Leukemia: A Consensus Document from the ELN-DAVID MRD Working Party. Blood. 2025 Dec 15:blood.2025031480. doi: 10.1182/blood.2025031480.

 CN
CN



