A research team from Jinan University (China), Emory University (USA), and Guangzhou Institutes of Biomedicine and Health, Chinese Academy of Sciences (China) published a study in Nature Biomedical Engineering (IF = 26.6) entitled Single-nucleus transcriptomics of an engineered pig model reveals microglia–T cell interactions driving Huntington’s disease neurodegeneration.
Using a genetically knock-in pig model of Huntington’s disease (HD), combined with single-nucleus transcriptomics, spatial transcriptomics, and immunological validation, the team systematically revealed for the first time the infiltration mechanism of CD8⁺ T cells in the HD striatum. They also elucidated the pathological process in which interferon-associated microglia recruit T cells via the CCL8–CCR5 chemotactic axis, which in turn induces neuronal damage through secretion of perforin and granzyme A.
iGeneTech was honored to provide a custom porcine immune repertoire detection kit for this study, supporting the construction of T-cell receptor (TCR) sequencing libraries in the HD-KI pig model.

Research Background: Mechanistic Mysteries and Model Limitations in Huntington’s Disease
Huntington’s disease is a fatal neurodegenerative disorder caused by abnormal expansion of CAG repeat sequences in the HTT gene. Its most prominent pathological feature is the selective loss of neurons in the striatum.
Although neuroinflammation is known to contribute to HD pathogenesis, previous studies have mainly focused on functional changes in microglia and astrocytes. Whether T cells infiltrate the brain parenchyma and their role in HD-specific neurodegeneration have long been controversial.
Existing mouse models of HD recapitulate some molecular pathological features but fail to reproduce the massive striatal neuronal loss seen in human HD patients. This limitation has severely hindered in-depth research into the mechanisms underlying selective neurodegeneration in HD.
Methodological Innovation: TCR Sequencing Identifies Clonal Expansion
The research team performed TCR β-chain sequencing on striatal and splenic tissues from HD-KI pigs using iGeneTech’s custom MultipSeq Panel. The results showed that infiltrating CD8⁺ T cells in the striatum exhibited markedly hyper-expanded TCR clonotypes, whereas the spleen was dominated by diverse clonotypes.
This finding demonstrates that brain-infiltrating T cells do not randomly cross the blood–brain barrier. Instead, they undergo antigen-driven clonal expansion in the periphery and are then specifically recruited to the striatum—elevating T cells from “bystanders” to key “executors” in HD pathogenesis.
Key Findings: CD8⁺ T Cells Drive Selective Neurodegeneration in HD
Through cross-species comparative analysis of single-nucleus transcriptomic data from HD-KI pigs, human HD patients, and HD mouse models, the team systematically uncovered the infiltration mechanism and neurotoxic role of CD8⁺ T cells in the HD striatum.
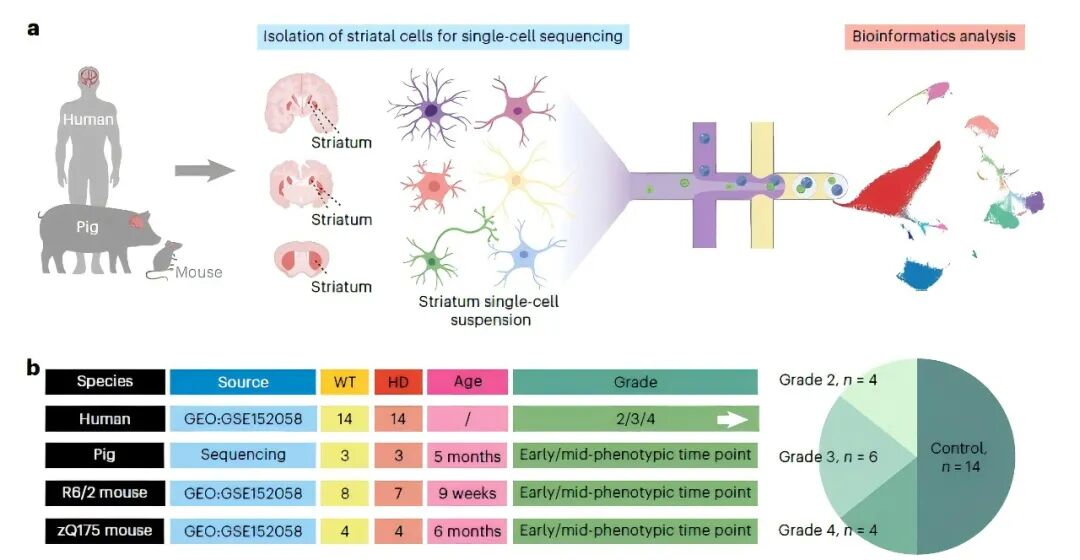
1. Cross-Species Comparison Reveals Specificity of T-Cell Infiltration
· The HD-KI pig model successfully recapitulated the selective loss of dSPNs and iSPNs in the striatum, as seen in human HD patients; no significant neuronal loss was observed in HD mice.
· Immunofluorescence confirmed a marked increase in CD8⁺ T cells in the striatum of both HD patients and HD-KI pigs, but complete absence in HD mice—indicating that mouse models cannot replicate T-cell-mediated immunopathology.
· TCR sequencing confirmed that striatal CD8⁺ T cells in HD-KI pigs were hyper-expanded clonotypes, distinct from the diverse clonal pattern in the spleen, proving they are antigen-driven “elite” effector cells.
2. Discovery of the Microglia–T Cell Interaction Axis
· The CCL8–CCR5 chemokine axis drives T-cell recruitment: CellPhoneDB analysis identified an IFN-associated microglial subset (MG2) that highly expresses CCL8, which specifically binds to the CCR5 receptor on CD8⁺ T cells.
· Spatial transcriptomics and in situ hybridization validated significant spatial colocalization of CCL8⁺ microglia and CCR5⁺ CD8⁺ T cells in the HD-KI pig striatum, confirming this axis mediates brain infiltration of T cells.
3. Neurotoxic Mechanism and Functional Validation
· The perforin–granzyme A pathway induces neuronal apoptosis: Infiltrating CD8⁺ T cells highly express PRF1 and GZMA, colocalize with neurons, and trigger apoptosis.
· In vitro experiments confirmed that treatment with perforin and granzyme A caused abnormal morphology and markedly increased TUNEL signals in primary neurons.
· CCL8 overexpression in HD mice successfully induced CD8⁺ T-cell infiltration, neuronal loss, and axonal demyelination—fully recapitulating the pathological features of the HD pig model.
·
Clinical Implications and Future Perspectives
· The extent of CD8⁺ T-cell infiltration and CCL8 expression levels in the striatum of HD patients are promising noninvasive predictive markers for assessing disease progression and neuronal injury.
· Tracking dynamic changes in dominant T-cell clones in peripheral blood using TCR sequencing may enable noninvasive monitoring of HD immunopathology, transforming current clinical management that relies mainly on motor symptoms and imaging.
·
Product Introduction
Based on its multiplex amplicon technology platform, iGeneTech has developed immune repertoire detection products covering humans and multiple model organisms (mouse, rat, pig, etc.). Leveraging the IMGT database, we offer custom panel services for multi-species immune repertoire research, providing personalized and integrated technical solutions for immune repertoire-related detection and scientific studies.
Product Information
Product Name | Specification | Cat. No. |
MultipSeq® Library Prep Kit V2 (100) | 16 rxn / 96 rxn | M61121 / M61122 |
IGT® UDI Primers 1-96 (10 μM each, for Illumina, plate) | 96 rxn | C80202 |
MultipSeq® Human TCR Research Assay (for Illumina) | 16 rxn / 96 rxn | M62021 / M62022 |
MultipSeq® Human BCR Research Assay (for Illumina) | 16 rxn / 96 rxn | M62041 / M62042 |
MultipSeq® Custom Panel | 16 rxn / 96 rxn | To be determined |
*Reagents for different sequencing platforms are available.

 CN
CN


