Fusion genes are the "golden targets" for precision tumor therapy. Different from ordinary gene mutations, a fusion gene is a chimeric gene formed by the recombination of partial or complete sequences of two different genes. Such chimeric genes will form abnormal transcripts or fusion proteins in subsequent biological processes, thereby inducing or promoting tumorigenesis, development and malignant transformation [1].
To date, more than 10,000 fusion genes have been found to be closely associated with tumorigenesis and development, and many of them are important components of tumor driver gene variations [2]. With the advancement of precision medicine, the detection of various fusion genes has been included in the routine recommendations of NCCN guidelines, such as for non-small cell lung cancer, colorectal cancer, hepatobiliary tumors and other malignancies [3,4].
Meanwhile, the types and indications of targeted drugs for fusion genes are constantly expanding. Taking non-small cell lung cancer (NSCLC) as an example, fusion genes such as ALK, ROS1, RET and NTRK are the most common clinical therapeutic targets, and the corresponding targeted drugs including crizotinib, alectinib and selpercatinib have been widely used in clinical practice. More notably, most of these drugs have been included in the national medical insurance catalog, which means the out-of-pocket expenses for patients are significantly reduced after medical insurance reimbursement, making precision therapy no longer a "luxury".
Note: All drugs are Category B medical insurance drugs, and the specific reimbursement ratio shall be subject to local medical insurance policies; some drugs were newly added to medical insurance in 2025 and will be implemented starting from January 1, 2026.
In the field of tumor precision diagnosis and treatment, fusion gene detection has always been a key link connecting diagnosis and treatment. How to ensure "no missed detection" in fusion gene testing?
Common molecular pathological detection methods for gene fusion currently include fluorescence in situ hybridization (FISH), reverse transcription polymerase chain reaction (RT-PCR), immunohistochemistry (IHC) and next-generation sequencing (NGS). Each detection method has distinct characteristics in clinical application, and a comprehensive judgment is often required by combining multiple detection methods in clinical practice. Compared with the three traditional single-gene fusion detection methods, NGS technology has received increasing attention in clinical testing because it can detect multiple genes at one time and sensitively and efficiently identify both known and unknown fusion forms [3,4].
DNA-based NGS is the foundation of tumor gene detection, with its core advantage being the simultaneous detection of multiple variation types such as single nucleotide variants (SNV), insertions and deletions (InDel), and copy number variants (CNV), providing comprehensive genomic information for clinical practice. However, detecting fusion genes at the DNA level has several intractable problems:
· Probes need to cover lengthy intronic regions with a large number of repetitive sequences;
· High GC content is not conducive to the capture of target region fragments by probes;
· Similar repetitive sequences in introns of different genes are not conducive to accurate sequence alignment, affecting detection accuracy;
· Complex transcriptional or post-transcriptional splicing processes may interfere with the identification of gene fusions.
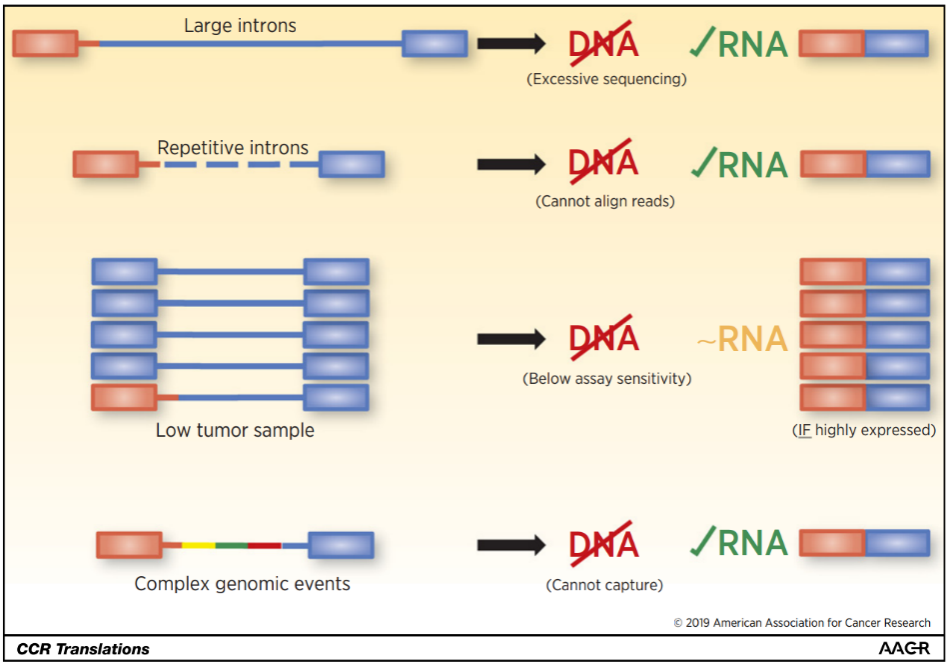
Figure 1. Schematic diagram of gene fusion [6]
In contrast to DNA detection, RNA detection directly targets spliced mRNA sequences, completely avoiding the interference of introns, accurately identifying clinically relevant fusion transcripts and significantly improving the detection rate of fusion genes [6].
Nevertheless, RNA detection also has obvious limitations: it has extremely high requirements for sample quality. Formalin-fixed paraffin-embedded (FFPE) samples are common clinical tumor samples; RNA degradation caused by prolonged storage or improper processing will directly affect the accuracy of RNA detection and even lead to missed detection.
How to Achieve More Accurate Detection?
Guidelines and consensus unanimously recommend: DNA+RNA NGS co-detection, integrating the advantages of both and making up for their respective limitations.
DNA and RNA detection have their own strengths and weaknesses and are highly complementary—neither can achieve a 100% fusion gene detection rate when used alone. Thus, the current optimal solution is DNA+RNA co-detection, which has been unanimously recommended by authoritative domestic and foreign guidelines and consensus.
· 2026 Version of Chinese Clinical Practice Expert Consensus on RNA-based Next-Generation Sequencing (NGS) for Fusion Gene Detection in Non-Small Cell Lung Cancer: Refines the application norms of RNA-NGS in NSCLC fusion gene detection, clarifies that RNA detection is a key means for accurate verification of fusion genes, and recommends its combined application with DNA detection.
· 2025 Version of Expert Consensus on Co-detection of Common Driver Genes in Solid Tumors by DNA and RNA High-Throughput Sequencing: Clarifies that combined detection by DNA-NGS and RNA-NGS can improve the fusion gene detection rate, standardizes the technical pathway of co-detection, and emphasizes its core position in the precision diagnosis and treatment of solid tumors.
· CSCO and NCCN Non-Small Cell Lung Cancer Guidelines: Mandatorily recommend DNA+RNA dual detection for populations with a high incidence of fusion genes (lung adenocarcinoma, female patients, never-smokers) to avoid missed detection; for patients with negative driver genes by DNA-NGS, supplementary RNA-NGS detection is prioritized to maximize the discovery of druggable targets.
How to select an efficient and reliable RNA detection technology has become one of the keys to clinical dual detection.
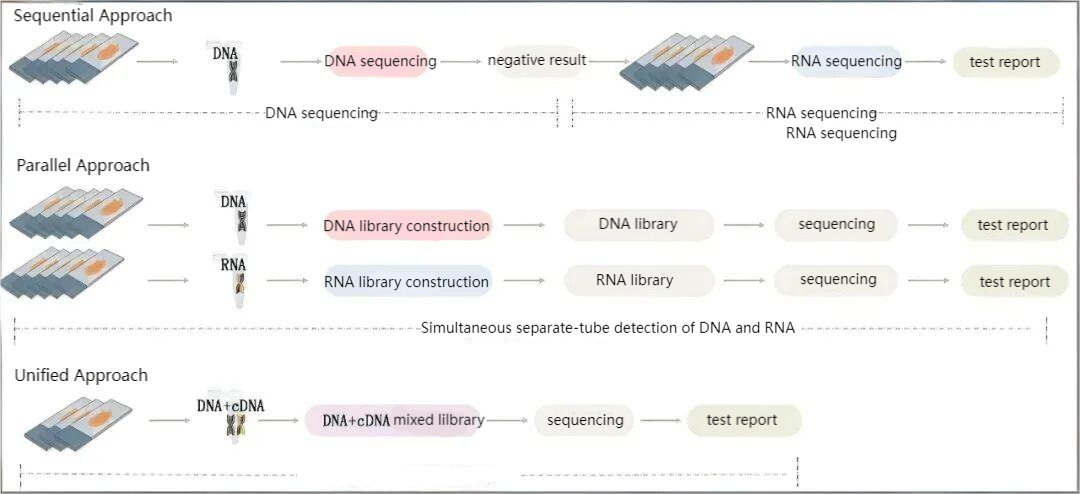
Figure 2. Schematic comparison of three strategies for DNA/RNA-NGS co-detection
DNA/RNA High-Throughput Sequencing Co-detection Strategies
Common capture technology schemes for DNA/RNA high-throughput sequencing detection include probe-based and amplicon-based methods. The probe-based method is suitable for sequential/parallel detection: probes are designed at the DNA level for SNV, InDel and introns of known fusions, and specific probes are designed at the RNA level for supplementary verification, with separate tube detection without mutual interference, making it applicable for multi-cancer detection. Although the amplicon-based method can achieve simultaneous DNA/RNA co-detection, it is limited by the detection region and the number of primers, and has strict requirements for FFPE sample quality, resulting in restricted clinical application.
The design principles for probes targeting fusion genes at the RNA level are as follows:
1. Conduct 3X tiled design based on all transcript sequences of the target gene, and remove redundant probes that repeatedly cover identical regions while ensuring 3X tiled coverage of each transcript.
2. Fully ensure the coverage of exon junction regions, and add additional probes at exon splicing regions during design to improve the detection rate of fusion genes.
3. For known fusions, especially low-frequency fusion variants, add mutant-type probes according to fusion sequences to improve the detection rate.

Figure 3. Probe design strategy for fusion gene detection at the RNA level
Currently, relying on 12 years of technical accumulation in targeted capture, iGeneTech has built a comprehensive RNA detection product system around the core principles of "precision, efficiency and compatibility". It aims to ensure multi-level precision, thereby solving the core pain points of RNA detection such as sample compatibility, interference elimination and accurate capture, and is applicable to a variety of clinical scenarios for DNA+RNA co-detection.
RNA Exome Whole Transcriptome Targeted Capture Product: An All-Round Capture Solution for Low-Quality Samples
Designed based on the RefGene/RefSeq database, it targets and captures a 46.1 Mb region, covering the coding regions of 20,761 genes and 69,209 transcripts as well as partial UTR regions. Its core features are:
· Precise fusion capture: Avoiding intron interference, probes are designed to match transcripts for efficient detection of fusion transcripts;
· Strong sample compatibility: Adapting to FFPE degraded samples, low-expression samples and samples with rare transcripts, solving sample quality pain points;
· High enrichment efficiency: Improving enrichment efficiency by more than 30% compared with whole transcriptome sequencing, reducing sequencing costs, and serving as a core tool for dual detection.

Figure 4. Capture performance test of RNA reference material.
Shuimu Jiheng RNA reference material was used to construct libraries at input amounts of 10 ng and 200 ng with an RNA library preparation kit, followed by hybrid capture and sequencing on an Illumina NovaSeq 6000 platform with PE150 mode.
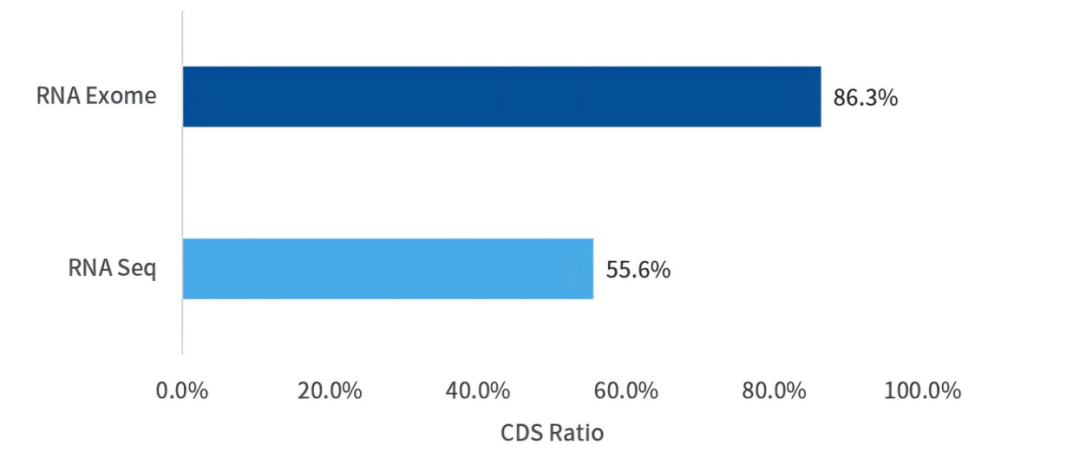
Figure 5. Proportion of sequencing data mapped to the CDS region using different technical protocols.
Total RNA input of 10 ng was analyzed by RNA‑Seq and RNA Exome respectively, and sequencing data were annotated against the CDS region.
Solid Tumor-Specific Fusion Gene Detection Panels: Precisely Matching Different Screening Needs
Adapted to multi-cancer detection needs and sensitive for fusion detection, with two types available:
· Comprehensive coverage panel: The 298 fusion gene panel covers known targets for multiple cancers, suitable for comprehensive screening of newly diagnosed patients and patients with unknown primary lesions.Note: Total RNA was extracted from Seraseq FFPE fusion RNA Reference Material v4 (Seraseq, cat. 10566485), and RNA libraries were constructed with an input amount of 50 ng. After capture, the libraries were sequenced and fusions were analyzed. The results showed that all fusion events were detected.
Table 1. Fusion results detected from positive fusion reference materials
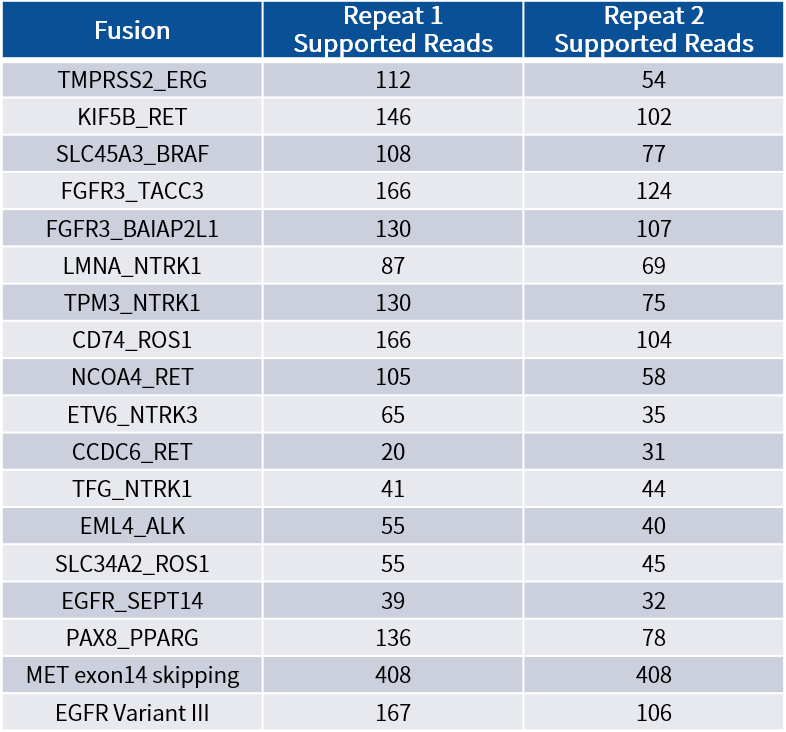
Note: Total RNA was extracted from Seraseq® FFPE Fusion RNA Reference Material v4 (Seraseq, Cat. No. 10566485). An RNA library was constructed with 50 ng input, followed by capture, sequencing, and fusion analysis. Results showed that all fusion events were detected.
· Cost-effective panel: The 90 core fusion gene panel focuses on clinically high-frequency targets, suitable for routine detection and medical insurance target screening, and is the first choice for primary hospitals.Note: Shuimu Jiheng RNA fusion reference material and Hela RNA sample were mixed, with a total input amount of 50 ng (fusion copy number about 100 copies) to construct RNA libraries. After capture, the libraries were sequenced and fusions were analyzed. The results showed that all fusion events were sensitively detected.
Table 2. Fusion results detected from positive fusion reference materials
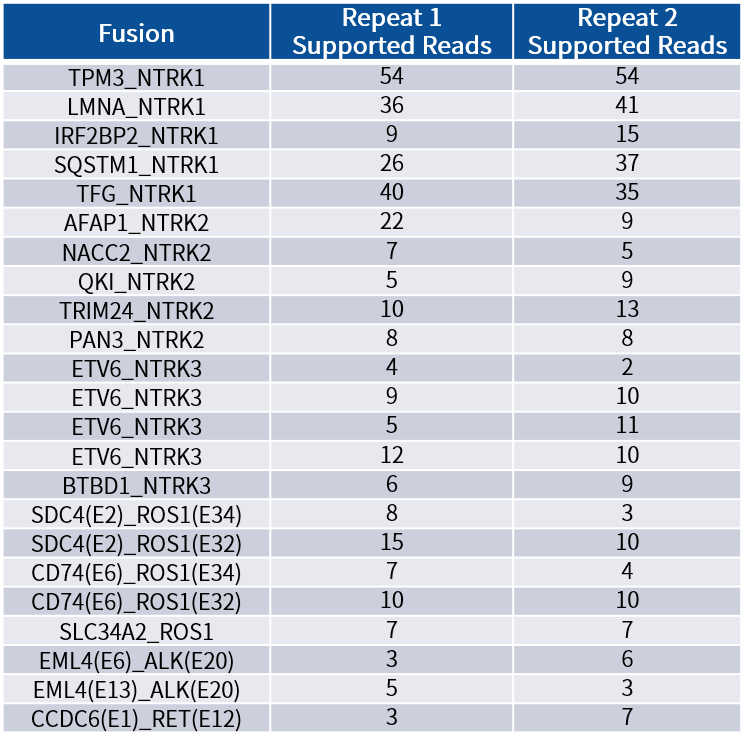
Note: Shuimu Jiheng RNA fusion reference material was mixed with Hela RNA sample at a total input of 50 ng, with approximately 100 fusion copies. An RNA library was constructed, followed by capture, sequencing, and fusion analysis. Results showed that all fusion events were sensitively detected.
Ultra-Fast Customization in 3–5 Days
Powered by the Ignite 3.0 high-throughput synthesis platform and independently developed massive parallel synthesis technology, iGeneTech can achieve full coverage and high uniformity of target sequences for Oligo Pools, with a single-run throughput of up to 680,000 independent sequences. Ultra-high throughput synthesis and rich design experience meet personalized needs and enable rapid customization of high-quality detection products.
In addition, iGeneTech also provides full-process supporting reagents for DNA+RNA detection as well as automated solutions, and can offer customized solutions according to clinical needs, helping to improve the precision, efficiency and accessibility of detection results.
Conclusion
Fusion gene detection is the first barrier to tumor precision therapy, and only accurate detection can safeguard patients' hope for targeted therapy. As an upstream kit developer for gene detection, iGeneTech has been deeply engaged in the field for 12 years, focusing on clinical needs, providing professional targeted capture solutions for clinical and scientific research institutions, and offering personalized customization services. We can customize exclusive detection solutions on demand, helping the standardized and large-scale implementation of DNA+RNA co-detection technology, enabling accurate detection to match targeted drugs, and jointly building a defense line for tumor precision diagnosis and treatment.
References
[1]. Gao Q, Liang WW, Foltz SM, Mutharasu G, Jayasinghe RG, Cao S, et al. Driver fusions and their implications in the development and treatment of human cancers. Cell Rep 2018;23:227–238.
[2]. Mertens F., Johansson B., Fioretos T., Mitelman F. The emerging complexity of gene fusions in cancer. Nat. Rev. Cancer. 2015;15:371–381. doi: 10.1038/nrc3905.
[3]. Xu C, Si L, Wang W, et al. Expert consensus on the diagnosis and treatment of NTRK gene fusion solid tumors in China. Thorac Cancer. 2022 Sep 20. doi: 10.1111/1759-7714.14644.
[4]. Ann Oncol. 2019 Sep 1;30(9):1417-1427. doi:10.1093/annonc/mdz204.
[5]. Schram, A. M. et al. Fusions in solid tumours: diagnostic strategies, targeted therapy, and acquired resistance. Nat. Rev. Clin. Oncol. doi:10.1038/nrclinonc.2017.127.
[6]. Kurtis D. Davies and Dara L. Aisne. Wake Up and Smell the Fusions: Single-Modality Molecular Testing Misses Drivers. Clinical Cancer Res 2019;25: 4586-4588.

 CN
CN








