Foreword
iGeneTech launches four syndrome pathogen identification panels covering 184 common pathogens. Paired with the IGT-AS01 automated workstation, the solution supports both manual and automated workflows, is compatible with diverse sample types, and requires no pre-cultivation or host depletion. It enables accurate pathogen identification and delivers an end-to-end solution for public health prevention, control, and research surveillance.
Background
Global public health prevention and control have drawn intense attention recently: Ethiopia has confirmed cases of Marburg virus disease, and Nipah virus outbreaks in Indonesia have triggered upgraded prevention measures in multiple countries. Meanwhile, the transition from winter to spring is a peak period for respiratory and gastrointestinal infectious diseases. Precise prevention and control of four major syndromes—fever with bleeding, respiratory, gastrointestinal, and encephalitis meningitis—have become key priorities in public health. Most of these syndromes are caused by various pathogens; some are highly pathogenic, lethal, and transmissible between humans. Rapid and accurate pathogen identification is critical for early detection and response in epidemic control.
The four major syndromes differ in clinical features, pathogenesis, and causative pathogens, yet all spread rapidly and carry risks of cluster infections or severe cases, posing challenges to public health and population health.

Respiratory Syndrome[1-4]:
Peak in autumn and winter, with common symptoms including fever, cough, nasal congestion, and sore throat, presenting as upper or lower respiratory tract infections. Major pathogens include influenza A virus, SARS-CoV-2, and Streptococcus pneumoniae. Transmission occurs mainly via droplets. High-risk groups include immunocompromised individuals and those with allergic constitutions, with frequent cluster infections in crowded settings.
Gastrointestinal Syndrome[5]:
A priority syndrome during winter–spring prevention and control, characterized by gastrointestinal disturbances such as nausea, vomiting, abdominal pain, and diarrhea. Common pathogens include norovirus, rotavirus, and Salmonella. Infections often result from contaminated food or environmental exposure. Mixed infections and emerging pathogens complicate clinical diagnosis and may lead to outbreaks.
Fever with Bleeding Syndrome[6-7]:
Defined by fever and systemic or localized bleeding. Typical pathogens include Crimean-Congo hemorrhagic fever virus, severe fever with thrombocytopenia syndrome virus (SFTSV), and dengue virus. After invasion, pathogens trigger a pathological cycle of inflammation activation, endothelial injury, and coagulation abnormalities. The syndrome is highly pathogenic and carries potential human-to-human transmission risk.
Encephalitis Meningitis Syndrome[8-9]:
A life-threatening syndrome caused by pathogens including Nipah virus, Japanese encephalitis virus, and Neisseria meningitidis. Pathogens cross the blood–brain barrier to invade the central nervous system, with typical symptoms of fever, headache, and altered consciousness. It progresses rapidly and has a high fatality rate.
Covered Species in the Identification Panels
| Respiratory Syndrome Pathogens | Gastrointestinal Syndrome Pathogens | Encephalitis Meningitis Syndrome Pathogens | Fever with Bleeding Syndrome Pathogens |
| Influenza A virus | Norovirus GI/GII | Japanese encephalitis virus | Dengue virus |
| Influenza B virus | Sapovirus | West Nile virus | Chikungunya virus |
| Influenza C virus | Rotavirus A/B/C | Dengue virus | Zika virus |
| Influenza D virus | Astrovirus | Zika virus | Ebola virus |
| Human parainfluenza virus 1 | Enteric adenovirus | Chikungunya virus | Marburg virus |
| Human parainfluenza virus 2 | Aichi virus | Nipah virus | Crimean-Congo hemorrhagic fever virus |
| Human parainfluenza virus 3 | Coxsackievirus | Enterovirus 71 | Rift Valley fever virus |
| Human parainfluenza virus 4 | Echovirus | Enterovirus | Lassa virus |
| Human respiratory syncytial virus A | Enterovirus 71 | Echovirus | Hantavirus |
| Human respiratory syncytial virus B | Hepatitis A virus | Coxsackievirus | SFTS virus |
| Human metapneumovirus | Hepatitis E virus | Varicella-zoster virus | Yellow fever virus |
| Human coronavirus 229E | Salmonella spp. | Cytomegalovirus | Kyasanur Forest disease virus |
| Human coronavirus OC43 | Shigella spp. | Epstein-Barr virus | Omsk hemorrhagic fever virus |
| Human coronavirus NL63 | Pathogenic Escherichia coli | Human herpesvirus 6 | Leptospira interrogans |
| Human coronavirus HKU1 | Campylobacter jejuni | Mumps virus | Rickettsia rickettsii |
| SARS-CoV | Yersinia enterocolitica | Measles virus | Orientia tsutsugamushi |
| SARS-CoV-2 | Vibrio cholerae | Rubella virus | Plasmodium falciparum |
| MERS-CoV | Vibrio parahaemolyticus | Rabies virus | Plasmodium vivax |
| Human adenovirus | Clostridium perfringens | Lymphocytic choriomeningitis virus | Plasmodium malariae |
| Human bocavirus | Clostridium difficile | Streptococcus pneumoniae | Plasmodium ovale |
| Rhinovirus A/B/C | Bacillus cereus | Neisseria meningitidis |
|
| Enterovirus | Listeria monocytogenes | Haemophilus influenzae |
|
| Epstein-Barr virus | Staphylococcus aureus | Listeria monocytogenes |
|
| Cytomegalovirus | Giardia lamblia | Escherichia coli K1 |
|
| Human herpesvirus 6 | Cryptosporidium spp. | Staphylococcus aureus |
|
| Human herpesvirus 7 | Cyclospora cayetanensis | Mycobacterium tuberculosis |
|
| Varicella-zoster virus | Entamoeba histolytica | Cryptococcus neoformans |
|
| Streptococcus pneumoniae |
| Cryptococcus gattii |
|
| Staphylococcus aureus |
| Aspergillus spp. |
|
| Haemophilus influenzae |
| Toxoplasma gondii |
|
| Legionella pneumophila |
| Naegleria fowleri |
|
| Mycoplasma pneumoniae |
| Acanthamoeba spp. |
|
| Chlamydia pneumoniae |
|
|
|
| Bordetella pertussis |
|
|
|
| Pseudomonas aeruginosa |
|
|
|
| Klebsiella pneumoniae |
|
|
|
| Acinetobacter baumannii |
|
|
|
| Moraxella catarrhalis |
|
|
|
| Pneumocystis jirovecii |
|
|
|
| Aspergillus fumigatus |
|
|
|
| Candida albicans |
|
|
|
Using multiple strain sequences from the NCBI database as references, iGeneTech designed pathogen identification panels with differentiated probe strategies: conserved within species, specific between species for bacteria, fungi, and parasites; full-genome coverage for viruses. This design ensures comprehensive, zero-miss detection of pathogens.
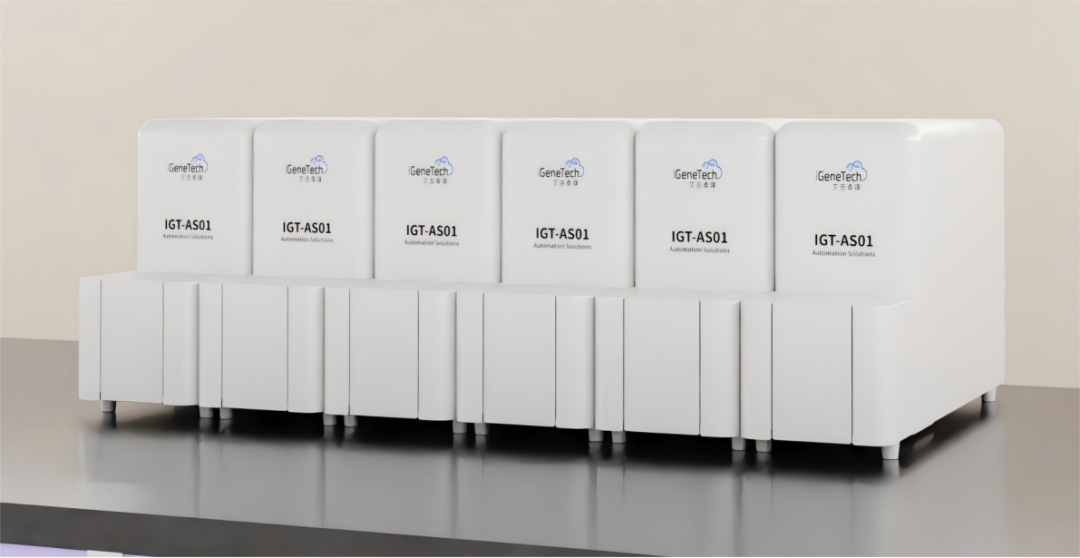

Paired with iGeneTech’s proprietary automated workstation reagent strip system, the workflow supports fast, independent processing of single samples and flexible throughput scaling via multi-unit arrays, greatly reducing manual handling. Only nucleic acid extracted from samples, capture probes, and index sequences need to be added to the reagent strips to start fully automated processing with no extra manual intervention, fully meeting laboratory demands for operational flexibility and minimal turnaround time.
Based on iGeneTech’s proprietary analytical pipeline for pathogen identification, raw sequencing data undergo quality control and host filtering; remaining reads are aligned against the pathogen reference database, and valid supporting read counts per species are tabulated and reported.
Key Advantages
Broad-spectrum and high-sensitivity, accurate and reliable
Covers diverse pathogens including viruses, bacteria, and fungi. Optimized probes stably capture trace pathogen nucleic acids. Combined with efficient hybridization and enrichment, the system delivers unbiased, precise genomic identification.
Simplified workflow, efficient and convenient
Supports diverse samples (wastewater, human, environmental, etc.). No pre-cultivation or host depletion required. Paired with 0.5–1 hour rapid hybridization and automated processing, it drastically simplifies pre-treatment and shortens detection time.
Fully integrated, walk�away detection
All-in-one reagent strips integrate library preparation and capture. Single-sample independent operation enables sample�in–library�out processing. Multi-instrument matrix expansion supports flexible throughput.
Physical isolation to prevent contamination, compact for strict environments
Independent reaction units eliminate cross-contamination. The compact footprint (only 0.1 m²) fits directly into biological safety cabinets, meeting high-level containment requirements.
End-to-End Detection Coverage
The four syndrome identification probe kits, combined with magnetic bead–based extraction kits, RNA pathogen library construction & capture kits, and DNA pathogen library construction & capture kits, provide a seamless sample-to-data workflow. They support batch processing with automated liquid handling workstations and are compatible with multiple high-throughput sequencing platforms to suit diverse scenarios.
Related Product Matrix: End-to-End Support from Extraction to Detection
Product Name | Specification | Cat. No. |
Encephalitis Meningitis Syndrome Panel | 16 / 96 rxn | PH2014841 / PH2014842 |
Respiratory Syndrome Identification Panel | 16 / 96 rxn | PH2014801 / PH2014802 |
Gastrointestinal Syndrome Identification Panel | 16 / 96 rxn | PH2014811 / PH2014812 |
Fever with Bleeding Syndrome Panel | 16 / 96 rxn | PH2014821 / PH2014822 |
Magnetic Beads Based Pathogen DNA/RNA Co-Extraction Kit | 50 rxn | E10021 |
Magnetic Beads Based Pathogen DNA/RNA Co-Extraction Kit (Host Depletion) | 50 rxn | E20011 |
IGT® DNA Pathogen Microbial Library Prep & Capture Kit(Illumina) | 16 rxn | C11361 / C11431 |
IGT® RNA Pathogen Microbial Library Prep & Capture Kit(Illumina) | 16 rxn | C11371 / C11441 |
IGT-AS12 Automated Liquid Handling Workstation (Configuration 3) | Configuration 3 | Q91013 |
References
[1] Gambotto A, Barratt-Boyes SM, de Jong MD, Neumann G, Kawaoka Y. 2008. Human infection with highly pathogenic H5N1 influenza virus. Lancet 371:1464–1475.[2] Gupta A, Madhavan MV, Sehgal K, Nair N, Mahajan S, et al. 2020. Extrapulmonary manifestations of COVID-19. Nat. Med. 26:1017–1032.[3] Andre G, Converso TR, Politano WR, Ferraz LF, Ribeiro ML, Leite LC, Darrieux M. 2017. Role of Streptococcus pneumoniae proteins in evasion of complement-mediated immunity. Front Microbiol 8:224.[4] Kim TS, Braciale TJ. 2009. Respiratory dendritic cell subsets differ in their capacity to support the induction of virus-specific cytotoxic CD8+ T cell responses. PLOS ONE 4:e4204.[5] Pathan N, Woolfall K, Popa M, et al. Selective digestive tract decontamination to prevent Healthcare associated infections in critically ill children: the PICNIC Multicentre randomised pilot clinical trial. Sci Rep 2023;13:21668. doi:10.1038/s41598-023-46232-7.[6] Zhang, W. K., Yan, J. M., Chu, M., Li, B., Gu, X. L., Jiang, Z. Z., … Yu, X. J. (2025). Bunyavirus SFTSV nucleoprotein exploits TUFM-mediated mitophagy to impair antiviral innate immunity. Autophagy, 21(1), 102–119. https://doi.org/10.1080/15548627.2024.2393067.[7] Wang, P., Liu, L., Liu, A., et al. Structure of severe fever with thrombocytopenia syndrome virus L protein elucidates the mechanisms of viral transcription initiation. Nat Microbiol 5, 864–871 (2020). https://doi.org/10.1038/s41564-020-0712-2.[8] Linder KA, Malani PN. Meningococcal Meningitis. JAMA. 2019;321(10):1014. doi:10.1001/jama.2019.0772.[9] Sala, F.A., Ditter, K., Dybkov, O., et al. Structural basis of Nipah virus RNA synthesis. Nat Commun 16, 2261 (2025). https://doi.org/10.1038/s41467-025-57219-5.

 CN
CN